Population genomics of the arboreal alligator lizard, Abronia campbelli
Authors
Natalie Winters, Rebecca Meyer, Morgan Muell, and Jamie Oaks
Introduction
Abronia alligator lizards are arboreal reptiles spanning southern Mexico and Central America. Some Abronia species endemic to eastern Guatemala, such as Abronia campbelli and Abronia frosti, are classified as critically endangered (Ariano-Sánchez & Melendez, 2009); the expansion of farmland in this area has left many populations of A. campbelli isolated to single trees. Understanding how the isolation and fragmentation of populations can affect genetic diversity and structure of a species is critical for informing conservation and management efforts. The goal of this project was to investigate whether recent extreme isolation of the agricultural populations has left a signature in the lizard’s genomes.
Methods
Collection, Genomic Library Prep and Sequencing
Blood samples from 45 Abronia lizards located in various populations in Eastern Guatemala were preserved on FTA cards.
Assembly of Genomic Loci
After extracting DNA from the blood samples, we sequenced tens of thousands of regions of the lizards’ genomes associated with the restriction sites (Bayona-Vásquez et al.). To assemble and align the sequence data across the lizard samples, we used ipyrad (Eaton & Overcast, 2020).
Principal Component Analysis (PCA)
To visualize the genetic variation among the Abronia alligator lizards, we used a principal component analysis.
IQ-TREE
To estimate the phylogenetic relationships among the sampled Abronia lizards, we used a maximum-likelihood search in IQ-TREE (Wong et al., 2025).
Results
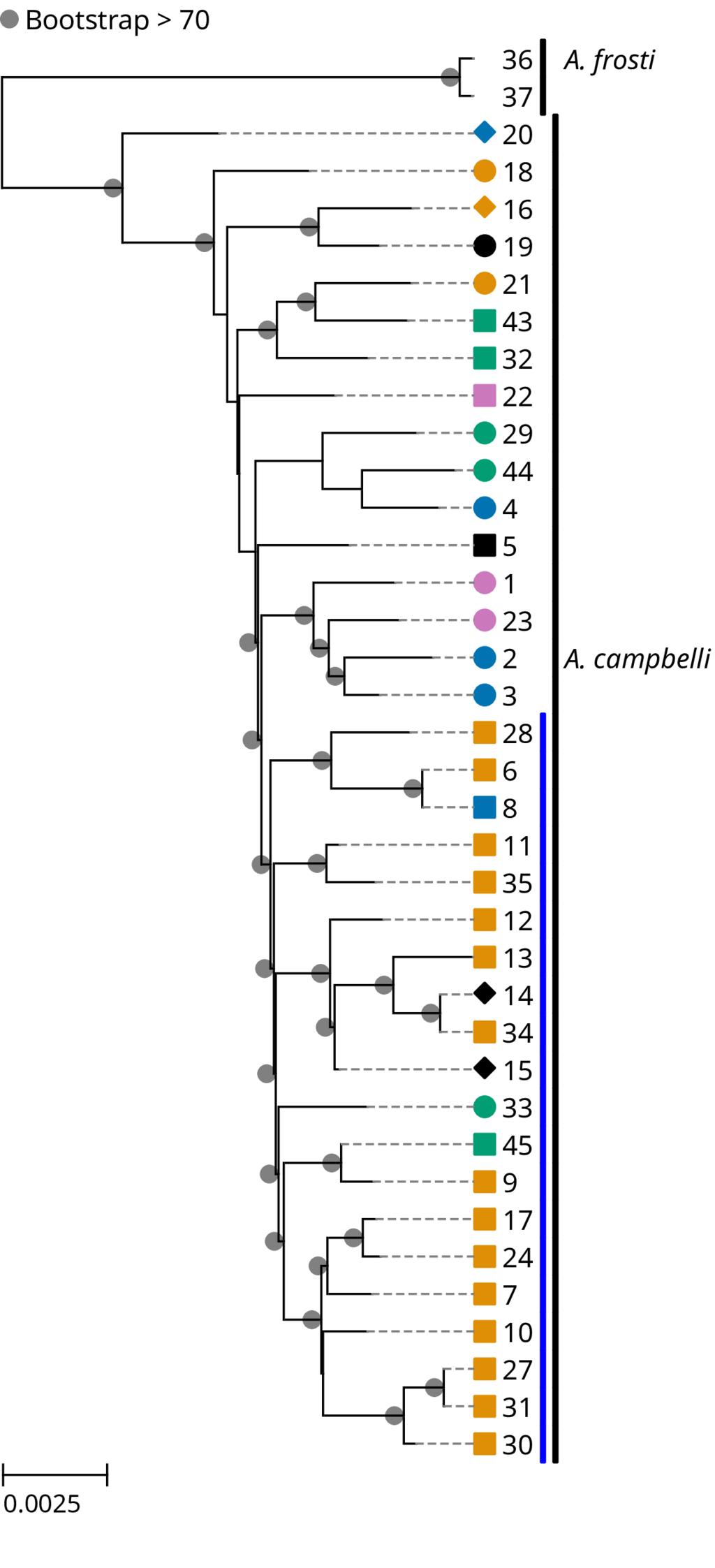
Figure 1. Estimated maximum-likelihood phylogenetic tree of the sampled *Abronia* lizards.
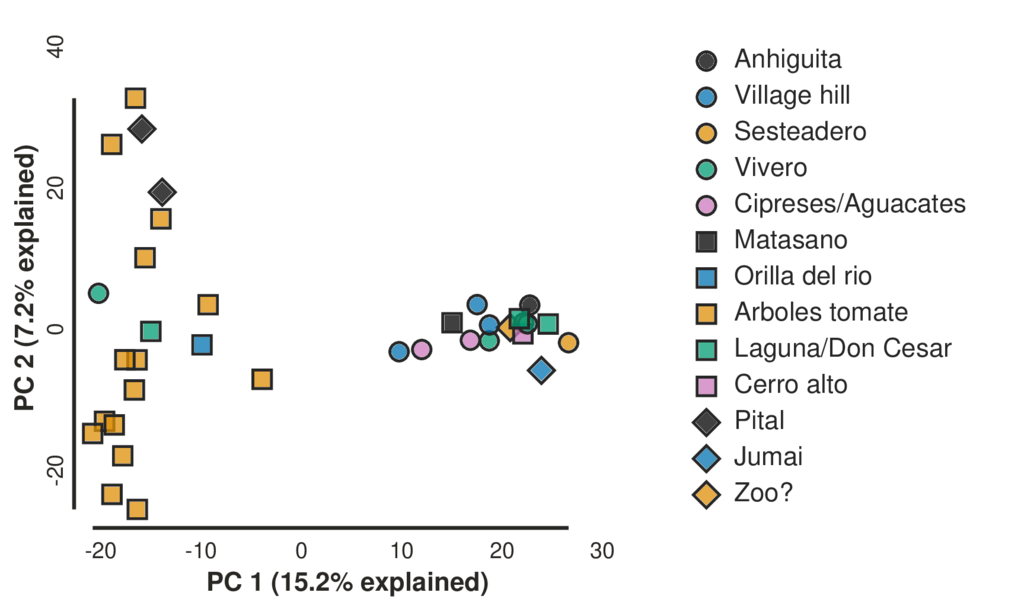
Figure 2. Projection of genetic variation along the first and second principal components (PC).
From the estimated phylogenetic tree (Figure 1), we can observe a monophyletic group of 20 individuals from five separate populations of various levels of isolation. The members of this clade group together along the first principal component in our PCA analysis (Figure 2).
This clade and group in the PCA does not correspond with any of the isolated agricultural locations from which the lizards were sampled. We do not see obvious signatures in the genomic diversity of the lizards left by recent extreme isolation
Conclusions
Within populations of Abronia campbelli in Eastern Guatemala, our principal component analysis, phylogenetic tree, and satellite data do not indicate a correspondence between extreme isolation imposed by agriculture and genomic divergence.
A better understanding of genetic variation among these isolated populations of Abronia in eastern Guatemala can be used to inform further conservation efforts and strategies.
References
- Wong TKF, Ly-Trong N, Ren H, Banos H, Roger AJ, Susko E, Bielow C, Maio ND, Goldman N, Hahn MW, Huttley G, Lanfear R, Minh BQ. 2025. IQ-TREE 3: Phylogenomic Inference Software using Complex Evolutionary Models. EcoEvoRxiv. DOI: 10.32942/X2P62N. Link.
- Eaton DAR, Overcast I. 2020. ipyrad: Interactive assembly and analysis of RADseq datasets. Bioinformatics 36:2592–2594. DOI: 10.1093/bioinformatics/btz966. Link.
- Ariano-Sánchez D, Melendez L. 2009. Arboreal Alligator Lizards in the Genus \textitAbronia: Emeralds from the Cloud Forests of Guatemala. Reptiles & Amphibians 16. DOI: 10.17161/randa.v16i1.15938. Link.
- Bayona-Vásquez NJ, Glenn TC, Kieran TJ, Pierson TW, Hoffberg SL, Scott PA, Bentley KE, Finger JW, Louha S, Troendle N, Diaz-Jaimes P, Mauricio R, Faircloth BC. Adapterama III: Quadruple-indexed, double/triple-enzyme RADseq libraries (2RAD/3RAD). PeerJ 7:e7724. DOI: 10.7717/peerj.7724. Link.
